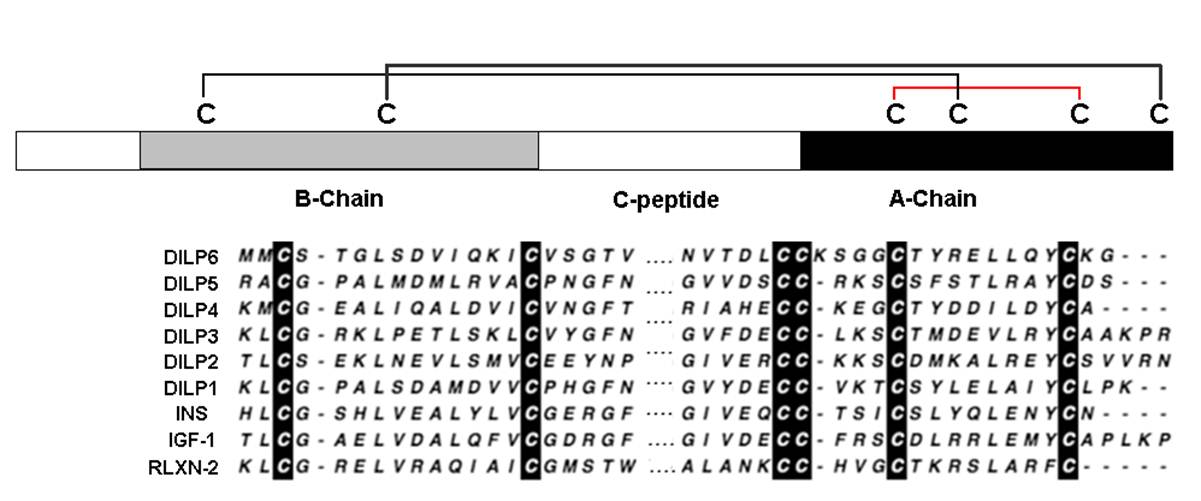
Insulin/IGF signaling (IIS) regulates essential processes including development, metabolism, and aging. The Drosophila genome encodes eight insulin/IGF-like peptide (dilp) paralogs, including tandem-encoded dilp1 and dilp2. Many reports show that longevity is increased by manipulations that decrease DILP2 levels. It has been shown that dilp1 is expressed primarily in pupal stages, but also during adult reproductive diapause. Here, we find that dilp1 is also highly expressed in adult dilp2 mutants under nondiapause conditions. The inverse expression of dilp1 and dilp2 suggests these genes interact to regulate aging. Here, we study dilp1 and dilp2 single and double mutants to describe epistatic and synergistic interactions affecting longevity, metabolism, and adipokinetic hormone (AKH), the functional homolog of glucagon. Mutants of dilp2 extend lifespan and increase Akh mRNA and protein in a dilp1-dependent manner. Loss of dilp1 alone has no impact on these traits, whereas transgene expression of dilp1 increases lifespan in dilp1 - dilp2 double mutants. On the other hand, dilp1 and dilp2 redundantly or synergistically interact to control circulating sugar, starvation resistance, and compensatory dilp5 expression. These interactions do not correlate with patterns for how dilp1 and dilp2 affect longevity and AKH. Thus, repression or loss of dilp2 slows aging because its depletion induces dilp1, which acts as a pro-longevity factor. Likewise, dilp2 regulates Akh through epistatic interaction with dilp1. Akh and glycogen affect aging in Caenorhabditis elegans and Drosophila. Our data suggest that dilp2 modulates lifespan in part by regulating Akh, and by repressing dilp1, which acts as a pro-longevity insulin-like peptide.Post S, Liao S, Yamamoto R, Veenstra JA, Nässel DR, Tatar M. Drosophila insulin-like peptide dilp1 increases lifespan and glucagon-like Akh expression epistatic to dilp2. Aging Cell. 2019;18(1):e12863.
Insulin and IGF signaling (IIS) is a complex system that controls diverse processes including growth, development, metabolism, stress responses, and aging. Drosophila melanogaster IIS is propagated by eight Drosophila insulin-like peptides (DILPs), homologs of both mammalian insulin and IGFs, with various spatiotemporal expression patterns and functions. DILPs 1-7 are thought to act through a single Drosophila insulin/IGF receptor, InR, but it is unclear how the DILPs thereby mediate a range of physiological phenotypes. We determined the distinct cell signaling effects of DILP2 and DILP5 stimulation upon Drosophila S2 cells. DILP2 and DILP5 induced similar transcriptional patterns but differed in signal transduction kinetics. DILP5 induced sustained phosphorylation of Akt, while DILP2 produced acute, transient Akt phosphorylation. Accordingly, we used phosphoproteomic analysis to identify distinct patterns of non-genomic signaling induced by DILP2 and DILP5. Across all treatments and replicates, 5,250 unique phosphopeptides were identified, representing 1,575 proteins. Among these peptides, DILP2, but not DILP5, dephosphorylated Ser15 on glycogen phosphorylase (GlyP), and DILP2, but not DILP5, was subsequently shown to repress enzymatic GlyP activity in S2 cells. The functional consequences of this difference were evaluated in adult Drosophila dilpmutants: dilp2 null adults have elevated GlyP enzymatic activity relative to wild type, while dilp5 mutants have reduced GlyP activity. In flies with intact insulin genes, GlyP overexpression extended lifespan in a Ser15 phosphorylation-dependent manner. In dilp2 mutants, that are otherwise long-lived, longevity was repressed by expression of phosphonull GlyP that is enzymatically inactive. Overall, DILP2, unlike DILP5, signals to affect longevity in part through its control of phosphorylation to deactivate glycogen phosphorylase, a central modulator of glycogen storage and gluconeogenesis.Post S, Karashchuk G, Wade JD, Sajid W, De meyts P, Tatar M. Insulin-Like Peptides DILP2 and DILP5 Differentially Stimulate Cell Signaling and Glycogen Phosphorylase to Regulate Longevity. Front Endocrinol (Lausanne). 2018;9:245.
How different organs in the body sense growth perturbations in distant tissues to coordinate their size during development is poorly understood. Here we mutate an invertebrate orphan relaxin receptor gene, the Drosophila Leucine-rich repeat-containing G protein-coupled receptor 3 (Lgr3), and find body asymmetries similar to those found in insulin-like peptide 8 (dilp8) mutants, which fail to coordinate growth with developmental timing. Indeed, mutation or RNA intereference (RNAi) against Lgr3 suppresses the delay in pupariation induced by imaginal disc growth perturbation or ectopic Dilp8 expression. By tagging endogenous Lgr3 and performing cell type-specific RNAi, we map this Lgr3 activity to a new subset of CNS neurons, four of which are a pair of bilateral pars intercerebralis Lgr3-positive (PIL) neurons that respond specifically to ectopic Dilp8 by increasing cAMP-dependent signalling. Our work sheds new light on the function and evolution of relaxin receptors and reveals a novel neuroendocrine circuit responsive to growth aberrations.This publication used Phoenix's DILP 8 (CAT# 035-79) for ligand-receptor interactions study.
Garelli A, Heredia F, Casimiro AP, et al. Dilp8 requires the neuronal relaxin receptor Lgr3 to couple growth to developmental timing. Nat Commun. 2015;6:8732.
Reduced insulin/IGF signaling extends lifespan in diverse species, including Drosophila melanogaster where the genome encodes seven insulin-like peptides (dilp1-7). Of these, reduced dilp2 expressed in the brain has been associated with longevity assurance when over-expression of dfoxo in fat bodies extends lifespan. Here, we show that the insulin-regulated transcription factor dFOXO positively modulates dilp6 mRNA in adult fat body. Over-expression of dilp6 in adult fat body extends lifespan and increases longevity-associated metabolic phenotypes. Adult fat body dilp6 expression represses dilp2 and dilp5 mRNA in the brain, and the secretion of DILP2 into the hemolymph. The longevity benefit of expressing dfoxo in fat body, and the nonautonomous effect of fat body dfoxo upon brain dilp expression, is blocked by simultaneously repressing dilp6 by RNAi in fat body. dilp6 thus appears to bridge dFOXO, adipose tissue and brain endocrine function to regulate Drosophila longevity.
Bai H, Kang P, Tatar M. Drosophila insulin-like peptide-6 (dilp6) expression from fat body extends lifespan and represses secretion of Drosophila insulin-like peptide-2 from the brain. Aging Cell. 2012;11(6):978-85.
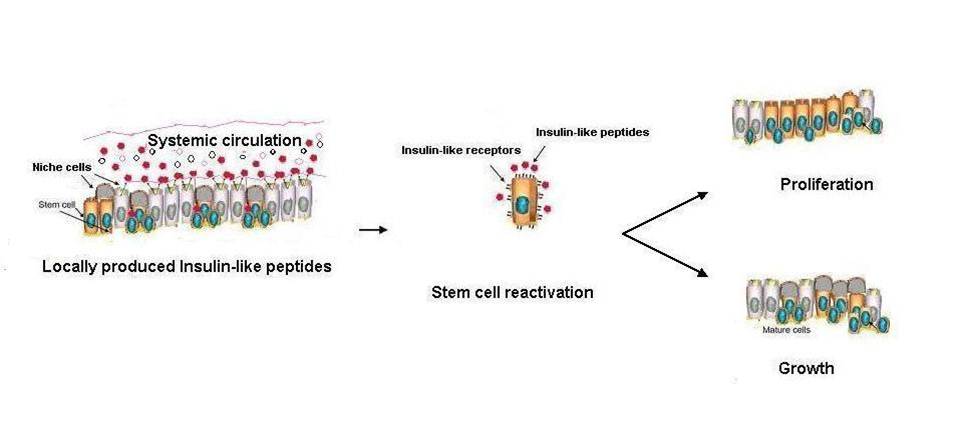
Many stem, progenitor and cancer cells undergo periods of mitotic quiescence from which they can be reactivated. The signals triggering entry into and exit from this reversible dormant state are not well understood. In the developing Drosophila central nervous system, multipotent self-renewing progenitors called neuroblasts undergo quiescence in a stereotypical spatiotemporal pattern. Entry into quiescence is regulated by Hox proteins and an internal neuroblast timer. Exit from quiescence (reactivation) is subject to a nutritional checkpoint requiring dietary amino acids. Organ co-cultures also implicate an unidentified signal from an adipose/hepatic-like tissue called the fat body. Here we provide in vivo evidence that Slimfast amino-acid sensing and Target of rapamycin (TOR) signalling activate a fat-body-derived signal (FDS) required for neuroblast reactivation. Downstream of this signal, Insulin-like receptor signalling and the Phosphatidylinositol 3-kinase (PI3K)/TOR network are required in neuroblasts for exit from quiescence. We demonstrate that nutritionally regulated glial cells provide the source of Insulin-like peptides (ILPs) relevant for timely neuroblast reactivation but not for overall larval growth. Conversely, ILPs secreted into the haemolymph by median neurosecretory cells systemically control organismal size but do not reactivate neuroblasts. Drosophila thus contains two segregated ILP pools, one regulating proliferation within the central nervous system and the other controlling tissue growth systemically. Our findings support a model in which amino acids trigger the cell cycle re-entry of neural progenitors via a fat-body-glia-neuroblasts relay. This mechanism indicates that dietary nutrients and remote organs, as well as local niches, are key regulators of transitions in stem-cell behaviour.
Sousa-nunes R, Yee LL, Gould AP. Fat cells reactivate quiescent neuroblasts via TOR and glial insulin relays in Drosophila. Nature. 2011;471(7339):508-12.
In metazoans, tissue growth relies on the availability of nutrients--stored internally or obtained from the environment--and the resulting activation of insulin/IGF signaling (IIS). In Drosophila, growth is mediated by seven Drosophila insulin-like peptides (Dilps), acting through a canonical IIS pathway. During the larval period, animals feed and Dilps produced by the brain couple nutrient uptake with systemic growth. We show here that, during metamorphosis, when feeding stops, a specific DILP (Dilp6) is produced by the fat body and relays the growth signal. Expression of DILP6 during pupal development is controlled by the steroid hormone ecdysone. Remarkably, DILP6 expression is also induced upon starvation, and both its developmental and environmental expression require the Drosophila FoxO transcription factor. This study reveals a specific class of ILPs induced upon metabolic stress that promotes growth in conditions of nutritional deprivation or following developmentally induced cessation of feeding.
Slaidina M, Delanoue R, Gronke S, Partridge L, Léopold P. A Drosophila insulin-like peptide promotes growth during nonfeeding states. Dev Cell. 2009;17(6):874-84.
Members of the insulin family of peptides have conserved roles in the regulation of growth and metabolism in a wide variety of metazoans. Here we show that Drosophila insulin-like peptide 6 (DILP6), which is structurally similar to vertebrate insulin-like growth factor (IGF), is predominantly expressed in the fat body, a functional equivalent of the vertebrate liver and adipocytes. This expression occurs during the postfeeding stage under the direct regulation of ecdysteroid. We further reveal that dilp6 mutants show growth defects during the postfeeding stage, which results in reduced adult body size through a decrease in cell number. This phenotype is rescued by fat body-specific expression of dilp6. These data indicate that DILP6 is a functional, as well as a structural, counterpart of vertebrate IGFs. Our data provide in vivo evidence for a role of ILPs in determining adult body size through the regulation of postfeeding growth.
Okamoto N, Yamanaka N, Yagi Y, et al. A fat body-derived IGF-like peptide regulates postfeeding growth in Drosophila. Dev Cell. 2009;17(6):885-91.
| Catalog# | Product | Standard Size | Price |
|---|---|---|---|
| 035-79 | Drosophila Insulin-Like Peptide 8 (DILP 8) (Drosophila melanogaster) | 100 µg | $571 |
| 036-18 | Drosophila Insulin-Like Peptide 1 (DILP 1) (Drosophila melanogaster) | 100 µg | $541 |
| 036-17 | Drosophila Insulin-Like Peptide 2 (DILP 2) (Drosophila melanogaster) | 100 µg | $541 |
| 036-71 | Drosophila Insulin-Like Peptide 3 (DILP 3) (A29/B25) (Drosophila melanogaster) | 100 µg | $541 |
| 035-97 | Drosophila Insulin-Like Peptide 3 (DILP 3) (Drosophila melanogaster) | 100 µg | $508 |
| 036-19 | Drosophila Insulin-Like Peptide 4 (DILP 4) (Drosophila melanogaster) | 100 µg | $481 |
| 035-96 | Drosophila Insulin-Like Peptide 5 (DILP 5), DB Variant (Drosophila melanogaster) | 100 µg | $508 |
| 035-98 | Drosophila Insulin-Like Peptide 6 (DILP 6) (Drosophila melanogaster) | 100 µg | $508 |
| 036-70 | Drosophila Insulin-Like Peptide 7 (DILP 7) (Drosophila melanogaster) | 100 µg | $662 |
Social Network Confirmation